When it comes to finding new chemical matter to engage biological targets, fragment-based screening (FBS) offers a number of advantages over traditional high-throughput screening (HTS). The success of this approach has resulted in a growing list of molecules advancing through clinical trials. From the perspective of a medicinal chemist, FBS makes it much faster to identify actionable hits as compared to plowing through the dross of a HTS campaign. The resulting leads tend to be of higher quality, exhibiting lower lipophilicity and lower molecular weight. The operational advantages of FBS make it particularly attractive, as a wider swath of chemical space can be searched more quickly, with fewer compounds, less protein consumption, and lower overall cost. Using the appropriate combination of biophysical and biochemical methods streamlines the validation process by removing possible artifacts at an early stage.
The availability of multiple screening methods is another consideration in choosing an FBS approach. As the most widespread approach, screening by NMR is a highly effective and versatile tool for identifying fragment hits. Even within the NMR paradigm, different detection methods (e.g. CPMG, STD, waterLOGSY) are susceptible to different artifacts such that the intersection of hit sets identifies true hits with high confidence. Other methods, such as MST, DSF and SPR; are commonly used both as primary screening tools and as downstream validation methods. Ideally, one chooses a workflow where initially identified hits are subsequently validated by a completely orthogonal technique. In my experience, the most successful embodiment is a NMR screen with follow-up by SPR.
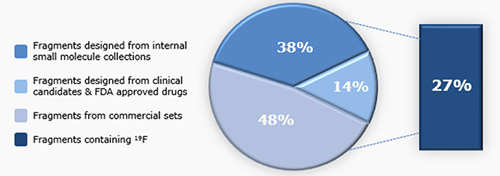
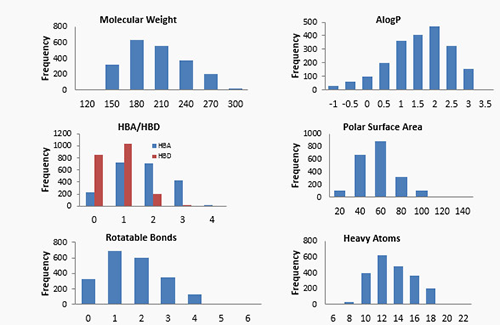
In applying FBS at ChemPartner, we provide discovery tools that can integrate with the existing strengths and capabilities of our partners. A key component of the FBS toolset is the fragment library. We built a fragment library using a threefold approach: (1) leveraging the expertise of our synthetic chemists to prepare uniquely designed fragments (2) extracting fragments from the structures of FDA approved drugs and (3) selecting fragments from commercially available sets. The resulting library offers a unique set of fragments, with overall property distributions amenable to finding high quality leads. We have successfully identified hits from this library for multiple target proteins, as illustrated by the case study with nicotinamide phosphoribosyltransferase (NAMPT).
As an essential cofactor required in all living cells, the availability of nicotinamide adenine dinucleotide (NAD) must be maintained for viability. NAD is an essential redox cofactor in cellular metabolism. NAD is also consumed as a substrate by enzymes associated with: DNA damage recognition and repair (e.g. PARP, calcium-mobilizing second messengers (e.g. cyclic ADP ribose synthetases), and transcriptional regulation (e.g. sirtuins). Cells obtain NAD by two pathways: de novo synthesis starting from amino acids, and a salvage pathway that recycles liberated nicotinamide. NAMPT is responsible for catalyzing the first of two steps in the salvage pathway [EC 2.4.2.12]. The wide-ranging physiological roles of NAD metabolism present numerous therapeutic opportunities across a range of indications. In particular, cancer cells show elevated sensitivity to the disruption of the salvage pathway.
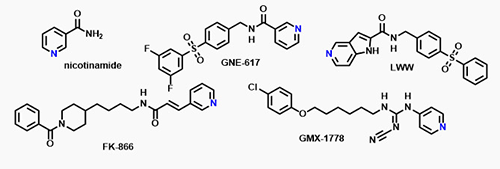
Early small-molecule NAMPT inhibitors derive inspiration from nicotinamide itself. The nucleophilic nitrogen in each of these substrate inhibitors is phosphorylated by NAMPT. While effective both in vitro and in animal tumor models, human clinical trials of FK866 and GMX1778 proved disappointing. Poor and variable oral bioavailability coupled with short plasma half-life of FK866 and GMX1777 necessitated administration by intravenous infusion. Lack of efficacy was hypothesized to result from insufficient target engagement prior to reaching toxicity inducing doses [6]. Subsequent work identified potent inhibition of CYP2C9 as a persistent problem with molecules bearing the substrate-like pyridyl nucleophile.
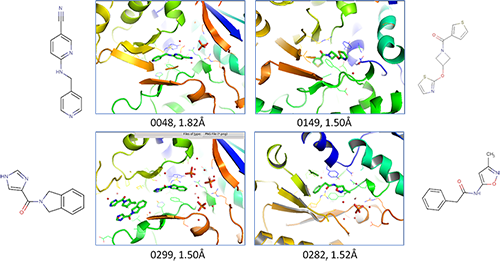
NAMPT was subjected to FBS by NMR with a one-thousand-member subset selected from the larger ChemPartner fragment library. After deconvolution and validation, more than 20 novel binding fragments were identified. Subsequently obtained X-ray crystal structures revealed some fragments bound in the active site, while other fragments bound to different regions of the protein. Specifically, compound 0048 shares the common nucleophilic pyridine motif present in nicotinamide and binds in a similar orientation. By contrast, fragments 0149, 0299 and 0282 lack this motif and bind elsewhere. Computational modeling suggests that hydrogen-bonding and pi-stacking interactions in these regions explain the fragment-protein interaction. The unique binding mode and chemical structure of each fragment illustrate how the FBS approach can quickly generate new starting points for novel lead identification.





